Single-cell & Single-nuclei RNAseq
From fresh and fixed tissue dissociation through Cellranger processing and downstream analysis, we handle the full single-cell workflow.
Why work with 3DG for Single-cell?
Single-cell projects fail more often at the sample prep stage than any other. We bring hands-on expertise in tissue dissociation, nuclei isolation, protein tagging, custom gene tracking – the steps that determine whether your experiment yields the best quality data. Every project gets direct scientific oversight, not cookie cutter execution from a protocol sheet.
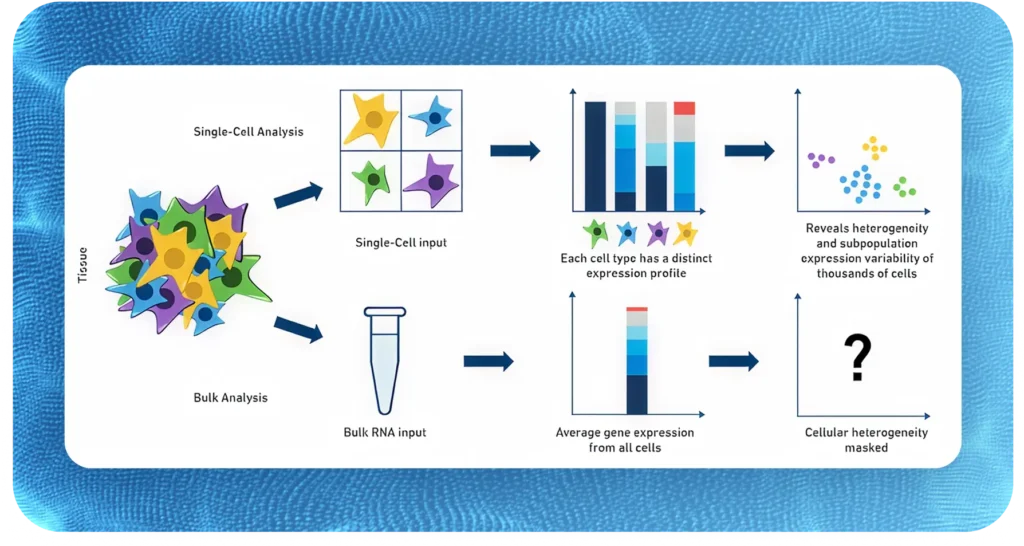
Single-cell or Single-nuclei?

Single-cell RNAseq
- Single-cell RNAseq (scRNAseq) uses intact, whole cells and captures the full cytoplasmic and nuclear transcriptome. It’s ideal when you can obtain high-quality, viable cells, such as from fresh tissues, blood, or cultured cells. You get a richer, more complete expression profile – but requires careful handling to preserve cell integrity.
- Use single cells when you want the fullest possible transcriptome and have access to fresh, viable samples such as cell culture cells or readily dissociated tissue.
Single-nuclei RNAseq
- Single-nuclei RNAseq (snRNA-seq) isolates just the nuclei and profiles nuclear RNA. It’s often used when intact cells are hard to obtain – like from frozen, archived, or tough tissues (e.g., brain, fibrotic tissue). It also reduces biases from dissociation stress (which can alter cell states) and can be a better choice when dealing with fragile or complex samples.
- Use single nuclei when working with frozen, delicate, or hard-to-dissociate tissues – or when minimizing stress artifacts is critical.
Fresh vs. Fixed Single-Cell
Single-cell platforms fall into two fundamentally different chemistries, and the distinction matters for experimental design, species compatibility, and which downstream assays are available.
Fresh cell capture (3′ and 5′ gene expression)
Fresh cell capture isolates intact, viable cells or nuclei from fresh or fresh-frozen tissue and captures mRNA via the poly-A tail. Because capture relies on polyadenylation rather than probe hybridization, this approach is species-agnostic and works on any organism that produces polyadenylated mRNA. It is the default chemistry for most single-cell experiments and supports the full range of multiomic add-ons: CITE-seq surface protein profiling, TCR/BCR immune repertoire sequencing, CRISPR perturbation screens, and ATAC-seq chromatin accessibility via the Multiome kit.
Fresh single-cell requires viable cells at the time of capture. Tissue dissociation quality is the primary determinant of data quality — poorly dissociated tissue, high ambient RNA from lysed cells, or low viability all degrade the resulting dataset. 3DG performs tissue dissociation in-house using the Miltenyi Octo dissociator with optimized protocols for diverse tissue types, including skin, tumor, and immune tissue.
Fixed cell capture (Chromium Fixed RNA, GEM-X Flex)
Fixed cell capture uses probe hybridization to capture gene expression from formalin-fixed cells or nuclei. This enables processing of FFPE tissue and allows samples to be fixed at the point of collection and batched for later processing, reducing technical variation across samples collected at different times or sites. It is the preferred approach for clinical sample cohorts where same-day processing is not feasible.
Because detection relies on pre-designed probe sets, fixed cell capture is currently limited to human and mouse. Custom probe design is available for genes outside the standard panel. Fixed chemistry supports multiplexing of up to 16 samples per GEM-X Flex reaction, substantially reducing per-sample reagent cost for large cohorts.
Single-nucleus RNA sequencing (snRNA-seq)
Single-nucleus RNA-seq isolates nuclei rather than intact cells, profiling nuclear RNA rather than the full cytoplasmic and nuclear transcriptome. It is the method of choice for tissues that are difficult or impossible to dissociate into viable single cells — brain, heart, skeletal muscle, kidney, and most frozen or FFPE archival material. Nuclei are more robust to mechanical and enzymatic dissociation and can be isolated from fresh-frozen or FFPE tissue sections.
The tradeoff is that nuclear RNA captures a different transcriptional signal than cytoplasmic RNA. Cytoplasmic transcripts are more abundant and provide a richer expression profile. Nuclear RNA is enriched for nascent transcripts and pre-mRNA, which can actually be informative for trajectory and RNA velocity analyses. For most cell type identification and differential expression work, snRNA-seq and scRNA-seq produce comparable results when analyzed with appropriate reference atlases.
Assay Variations available at 3DG
Standard single-cell RNA-seq measures gene expression. A growing set of assay variations captures additional molecular layers from the same cells simultaneously, linking transcriptional state to chromatin accessibility, surface protein abundance, immune receptor sequences, or genetic perturbations. Each add-on answers a different biological question and involves specific reagent, sample, and computational requirements.
Multiome: RNA + chromatin accessibility
The 10x Genomics Multiome kit captures gene expression and ATAC-seq chromatin accessibility from the same nucleus, enabling direct linkage of open chromatin regions to gene expression at single-cell resolution. This is the most informative single assay for understanding gene regulatory programs — paired RNA and ATAC data supports SCENIC+ regulatory network inference, peak-to-gene linkage, and transcription factor binding site validation that is not possible from RNA alone.
Multiome requires nuclei isolation rather than intact cells, making it compatible with fresh-frozen tissue and some fixed-frozen samples. It is not currently compatible with FFPE material. Sequencing requirements are approximately double that of standard scRNA-seq since two libraries are generated per sample. Computational analysis uses ArchR for the ATAC component and Seurat WNN or equivalent for joint embedding.
CITE-seq: surface protein profiling
CITE-seq (Cellular Indexing of Transcriptomes and Epitopes by Sequencing) measures surface protein abundance alongside gene expression using antibody-derived tags (ADTs). Antibodies conjugated to DNA barcodes bind to surface proteins; the barcodes are captured and sequenced alongside the transcriptome, providing simultaneous protein and RNA measurements from each cell.
CITE-seq is particularly valuable in immune profiling, where canonical surface markers such as CD4, CD8, CD19, and CD56 provide cleaner cell type discrimination than transcriptomics alone, and where protein-level information connects single-cell data to flow cytometry or mass cytometry results from the same study. It is also useful when the research question specifically involves surface receptor expression, ligand binding, or protein-level validation of transcriptional findings.
3DG uses the 10x Genomics Feature Barcode technology for CITE-seq. Panel design — selecting which antibodies to include — requires consideration of the cell types expected in the sample, antibody titration, and potential cross-reactivity. We work with clients to select appropriate panels from validated commercial sources (BioLegend TotalSeq) or can assist with custom panel design.
Immune profiling: TCR and BCR repertoire sequencing
V(D)J sequencing from the same cells as gene expression data links T cell and B cell receptor sequences to transcriptional state, enabling direct assignment of clonotype identity to cell type and functional annotation. This is essential for studies of adaptive immune responses, autoimmunity, CAR-T cell products, and tumor-infiltrating lymphocyte characterization.
TCR sequencing identifies alpha/beta (and gamma/delta) T cell receptor sequences and assigns clonotype identities. BCR sequencing identifies immunoglobulin heavy and light chain sequences and can track B cell somatic hypermutation and clonal expansion. Both are available as add-ons to standard 3′ or 5′ gene expression libraries using the 10x Genomics Immune Profiling kit. 5′ chemistry is required for V(D)J, it is not compatible with 3′ capture. Immune profiling is discussed in the context of 5′ end capture on the 10X Genomics support site.
Clonotype data is integrated with gene expression clustering to identify which transcriptional states are associated with clonal expansion, exhaustion, or antigen-specific activation. This integration is standard in the Cell Ranger VDJ pipeline and downstream tools including Scirpy (Python) and immunarch (R).
CRISPR screening
Single-cell CRISPR screens combine pooled genetic perturbations with single-cell transcriptomics to measure the transcriptional consequence of each perturbation at single-cell resolution. Each cell receives one or more guide RNAs targeting a gene of interest; the guide RNA identity is captured alongside the transcriptome, enabling direct comparison of perturbed and unperturbed cell states across thousands of perturbations in a single experiment.
This approach — variously called Perturb-seq, CROP-seq, or CRISP-seq depending on the implementation — is used to map gene function at transcriptome-wide scale, identify genetic interactions, and validate drug targets. 3DG supports CRISPR screening experiments using the 10x Genomics CRISPR Screening feature barcode workflow. Guide library design, lentiviral delivery, and selection are typically performed by the client lab; 3DG handles single-cell capture, library preparation, sequencing, and computational analysis of the resulting perturbation data.
Custom gene expression panels
The standard 10x Genomics whole-transcriptome assays capture all polyadenylated mRNA in each cell. For some applications, targeted gene expression panels offer advantages: lower sequencing cost per cell, higher sensitivity for low-abundance transcripts, and the ability to include non-polyadenylated targets such as certain non-coding RNAs or pathogen transcripts.
The 10x Genomics Fixed RNA Profiling kit (GEM-X Flex) supports custom probe design for targets outside the standard human and mouse panels. Custom panels can include virus transcripts, transgenes, reporter constructs, or species not covered by standard probesets. Panel design requires probe design and synthesis lead time of approximately 8-10 weeks and a minimum order quantity; 3DG can advise on feasibility and lead time for specific targets.
Multiplexing
Sample multiplexing allows multiple biological samples to be pooled into a single 10x reaction, reducing batch effects and reagent costs for large studies. Several multiplexing strategies are available depending on the sample type and assay chemistry.
Cell hashing (HTO multiplexing) uses antibody-derived hashtag oligos to label cells from different samples before pooling. Samples are demultiplexed computationally after sequencing based on hashtag signal. This approach works with any fresh cell sample and is compatible with 3′ and 5′ gene expression, CITE-seq, and V(D)J assays.
The GEM-X Flex fixed RNA kit supports up to 16-sample multiplexing natively using probe-based barcoding without antibodies, making it the preferred approach for clinical cohort studies where samples are collected over time and batched for processing. 10x Genomics CellPlex offers a reagent-based multiplexing option for fresh cells with up to 12 samples per reaction.
Our Process
What you provide
Tissue samples or single-cell suspensions
Single-cell suspensions should be at a minimum concentration of 400 cells/ul with a viability greater than 50%, with at least 100K cells total per sample.
What 3D Genomics provides
- Cell preparation including tissue dissociation using the Miltenyi Octo dissociator, dead cell removal and/or nuclei isolation, if applicable.
- Single cell isolation, cDNA Synthesis and library preparation including QC for sequencing. 3D Genomics supports all 10x Genomics single cell assays and other platforms.
- Sequencing, read mapping and alignment using CellRanger as well as other pipelines and informatic tools.
What 3D Genomics delivers
- Raw sequencing files (FASTQ) and sequencing QC report.
- For 10x Assays the 10x Cell Ranger output: web summary, feature-barcode matrices, BAM files, automated secondary analyses, Loupe browser files and other output files
- Computational support including clustering, cell type annotation and multiomic integration, if applicable
- Remaining tissue samples will be discarded or returned, as per your instructions
Ready to move your spatial and single cell research forward?
Tell us about your samples and research questions. We’ll take it from there.


